Individual-Based Modeling of Ecological and Evolutionary Processes
The authors present a review of the general concepts and an overview of the uses of individual-based models (IBM) in ecology. Over the ~15 years (as of 2005), the use of individual based models (discovered by a search for the keywords “individual-based”/”individual-oriented” and “model”) has increased (from 1 to 150 papers; I did a simple scholar search including “ecology” and got 4,260 results). While the authors confer that a strict definition of an IBM does not exist, they present that an IBMs simulate populations of individual agents with each having specific, unique traits that vary between agents. These models are designed to capture the variation of individuals in the population; however, the included variation should be related to the question being addressed.
The included variation and the resolution of the model at the individual scale can provide insight to the dynamics of the system at multiple different scales: population, community, and ecosystem. While the authors recognize that the “classical model” (differential equations and difference equations) have been extended to include some individual variation however they posit that some phenomena require us to zoom into the individual level. They partition the traits of the individuals into 5 “axes”:
- Spatial variability, local interactions, movement: classical models do not take into account separate individuals which can create non-uniformity
- Life cycle detail: difference equation (age/stage structured) models do capture some key dynamics of the life cycle of organisms; individual level resolution becomes advantageous when a large number of subclasses.
- Phenotypic variation and behavior: classical models can have trouble when partitioning the differences between individuals in terms of their phenotype and past experiences (behavior) becomes large (according to the authors greater than 2)
- Experience and learning: classical models have included implicit learning however IBMs can create individual experiences that shape individuals.
- Genetic and evolution: IBMs allow for more flexibility than classical models for including genetic traits and mimic real populations.
The authors present a “framework” where each trait is an axis creating a 5 dimensional space. Increasing along each axis, the details of mechanisms increase. Classical models, as defined by the authors, is positioned at the center (origin). IBMs diverge from the origin along any number of the axes simultaneously.
After reviewing ~900 papers, the authors identify 7 major groups of studies and summarize some studies related to each: movement through space, formation of patterns among individuals, for foraging and bioenergetics to population dynamics, exploitative species interactions, local competition and community dynamics, evolutionary processes, management-related processes.
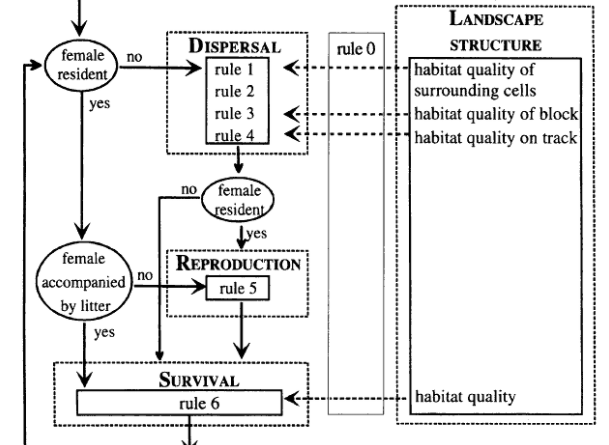
When developing IBMs, the state variables can be represented as table with each row is an individual and each column is their trait. Dynamics of the model are created when iterating over individuals. Other programs (such as NetLogo) are needed to apply these models fully. Below is an example of a flow diagram of a IBM describing a bear population (Wiegand et al. 1999).
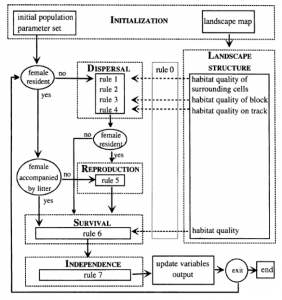
Individual-based models are a tool that can be used in models that transverse levels of organization (scales). Specifically, these models are at the resolution of the individual and can help us understand some phenomena at a higher scale (see “definition” the authors provide). Potentially, specific components/mechanisms of a system could be represented by IBMs where at a “higher” scale a variable is an output of the IBM.
DeAngelis, D. L., and W. M. Mooij. 2005. Individual-Based Modeling of Ecological and Evolutionary Processes. Annual Review of Ecology, Evolution, and Systematics 36:147–168. (